This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Future directions for the study of tubby and obesity
While the mechanism by which tubby controls metabolism and body weight is still unclear, several potential hypotheses currently exist and are discussed as they relate to the tubby protein pathway here. One of these hypotheses is that tubby normally regulates food intake to maintain proper energy homeostasis. This idea is supported by correlating three phenotypes previously observed in tubby mice, namely increased food intake, altered hunger neuropeptide expression, and obesity. [1,2] This role for TUB is also supported by the fact that TUB is the only TULP family member to be associated with obesity. One hypothesis for future testing is that TUB regulates hunger neuropeptides to control feeding behavior and energy homeostasis. Genomic and proteomic approaches would be very useful in an initial assessment of this hypothesis and preliminary results investigating this hypothesis are shown below.
Aim 1: Are there regions unique to Tub that could control feeding behavior?
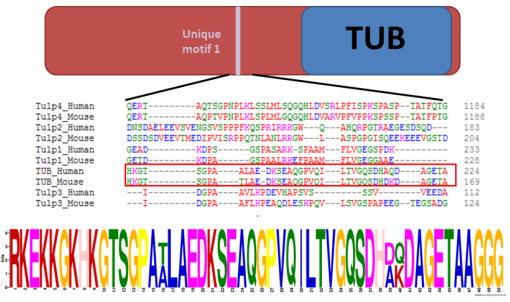
Since TUB is the only TULP family member to be associated with obesity, one could think that regions that are unique to TUB (not found in other TULPs) could be involved in its regulation of food intake and energy homeostasis. Alignments using Clustal Omega, Muscle, and T-Coffee were used to identify regions that were not conserved between TUB and other TULPs in mice and humans and two unique regions were identified. MEME was then used to identify predicted protein motifs within these regions (Figure 1 and Figure 2).
Future studies with these protein motifs would include mutating them within mice or other model organisms and determining whether this affects food intake. Since these regions are unique to TUB, a protein shown to affect food intake, I would predict that mutation of these motifs would result in increased food intake, and therefore, obesity.
Aim 2: Are there conserved DNA motifs in hunger neuropeptides where tubby could bind to regulate expression?
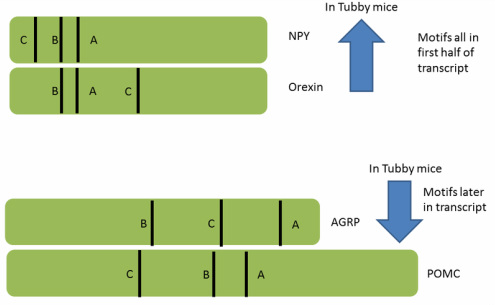
Since the hunger neuropeptides NPY, POMC, AGRP, and Orexin show altered expression in tubby mice, it is predicted that tubby normally regulates the expression of these neuropeptides, which in turn control feeding behavior. DREME was used to identify three conserved DNA motifs in these hunger neuropeptides. Interestingly, their relative location within the transcript correlated with whether they showed increased or decreased expression in tubby mice as shown in Figure 3. This finding strengthens the hypothesis that wild-type TUB may bind to these motifs to correctly regulate their expression, a process that is disrupted by the tubby mutation.
Future research could determine if TUB itself binds to these DNA motifs by performing Restriction Endonuclease Protection Selection and Amplification (REPSA) with the hunger neuropeptide DNA motifs A, B, and C above and TUB. [3] If TUB is found to bind and regulate these hunger neuropeptides it would provide further insight into tubby's regulation of feeding behavior and energy homeostasis.
Aim 3: DOES TUBBy affect food choice in addition to food intake?
A previous study by Vliet-Ostaptchouk et al. found that women carrying SNPs in Tub consumed more carbohydrates. [4] Similarly tubby mice also fail to activate their carbohydrate metabolism leading to a reliance on fat metabolism for energy needs. [5] Therefore another line of future research lies in determining whether TUB affects food choice in addition to food intake. To study this research question tubby mice would be given the choice between a high carbohydrate, high fat, and high protein diet and the amount they consume of each will be recorded.
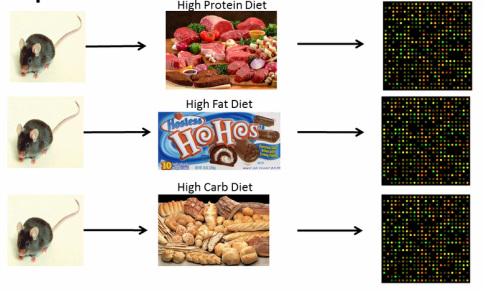
In order to determine what affect this food choice has on gene expression, wild-type mice will be assigned to either a high carbohydrate, high fat, or high protein diet and then microarray will be used to determine gene expression changes (Figure 4). Since TUB functions in energy metabolism, I would predict that genes associated with that GO term would show differential expression.
In order to determine what affect this food choice has on gene expression, wild-type mice will be assigned to either a high carbohydrate, high fat, or high protein diet and then microarray will be used to determine gene expression changes (Figure 4). Since TUB functions in energy metabolism, I would predict that genes associated with that GO term would show differential expression.
| future_directions_final_presentation.pdf | |
| File Size: | 1650 kb |
| File Type: | |
references
Cover Photo Credit
[1] Backberg, M., Madjid, N., Ogren, S.O., and Meister, B. (2004). Down-regulated expression of agouti-related protein (AGRP) mRNA in the hypothalamic arcuate nucleus of hyperphagic and obese tub/tub mice. Molecular Brain Research, 125 (129). doi:10.1016/j.molbrainres.2004.03.012.
[2] Guan, X.M., Yu, H., Lex, H.T., & Ploeg, V. (1998). Evidence of altered hypothalamic pro-opiomelancortin/neuropeptide Y mRNA expression in tubby mice. Molecular Brain Research, 59(273). doi: 10.1016/S0169328X98001508.
[3] Van Dyke, M.W., Van Dyke, N., and Sunavala-Dossabhoy, G. (2007). REPSA: General combinatorial approach for identifying preferred ligand–DNA binding sequences. Methods Related to DNA Sequence Recognition, 42 (2), 118. doi:10.1016/j.ymeth.2006.09.008
[4] vs Vliet-Ostaptchouk, J.V., Onland-Moret, N.C., Shiri-Sverdlov, R., et al. (2008). Polymorphisms of the TUB Gene Are Associated with Body Composition and Eating Behavior in Middle-Aged Women. PLoS ONE, 3(1), e1405. doi:10.1371/journal.pone.0001405.
[5] Wang, Y., Seburn, K., Bechtel, L., Lee, B.Y., Szatkiewicz, J.P., Nishina, P.M., and Naggert, J.K. (2006). Defective carbohydrate metabolism in mice homozygous for the tubby mutation. Physiol Genomics, 27(131). doi:10.1152/physiolgenomics.00239.2005.
[1] Backberg, M., Madjid, N., Ogren, S.O., and Meister, B. (2004). Down-regulated expression of agouti-related protein (AGRP) mRNA in the hypothalamic arcuate nucleus of hyperphagic and obese tub/tub mice. Molecular Brain Research, 125 (129). doi:10.1016/j.molbrainres.2004.03.012.
[2] Guan, X.M., Yu, H., Lex, H.T., & Ploeg, V. (1998). Evidence of altered hypothalamic pro-opiomelancortin/neuropeptide Y mRNA expression in tubby mice. Molecular Brain Research, 59(273). doi: 10.1016/S0169328X98001508.
[3] Van Dyke, M.W., Van Dyke, N., and Sunavala-Dossabhoy, G. (2007). REPSA: General combinatorial approach for identifying preferred ligand–DNA binding sequences. Methods Related to DNA Sequence Recognition, 42 (2), 118. doi:10.1016/j.ymeth.2006.09.008
[4] vs Vliet-Ostaptchouk, J.V., Onland-Moret, N.C., Shiri-Sverdlov, R., et al. (2008). Polymorphisms of the TUB Gene Are Associated with Body Composition and Eating Behavior in Middle-Aged Women. PLoS ONE, 3(1), e1405. doi:10.1371/journal.pone.0001405.
[5] Wang, Y., Seburn, K., Bechtel, L., Lee, B.Y., Szatkiewicz, J.P., Nishina, P.M., and Naggert, J.K. (2006). Defective carbohydrate metabolism in mice homozygous for the tubby mutation. Physiol Genomics, 27(131). doi:10.1152/physiolgenomics.00239.2005.
Site created by Rachael Baird.
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14