This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Protein interaction networks
A single protein, such as TUB, does not exist in isolation. Instead, it interacts with other proteins and molecules within the cell to modify function. Therefore, by studying the interaction partners of a protein, a better understanding of how that protein may function in the cell can be gained. The STRING database can be used to determine known and predicted protein interactions, these include direct, physical interactions and indirect associations inferred from genomic context, experiments, conserved coexpression, and data-mining in PubMed. [1]
Protein interactions of tubby
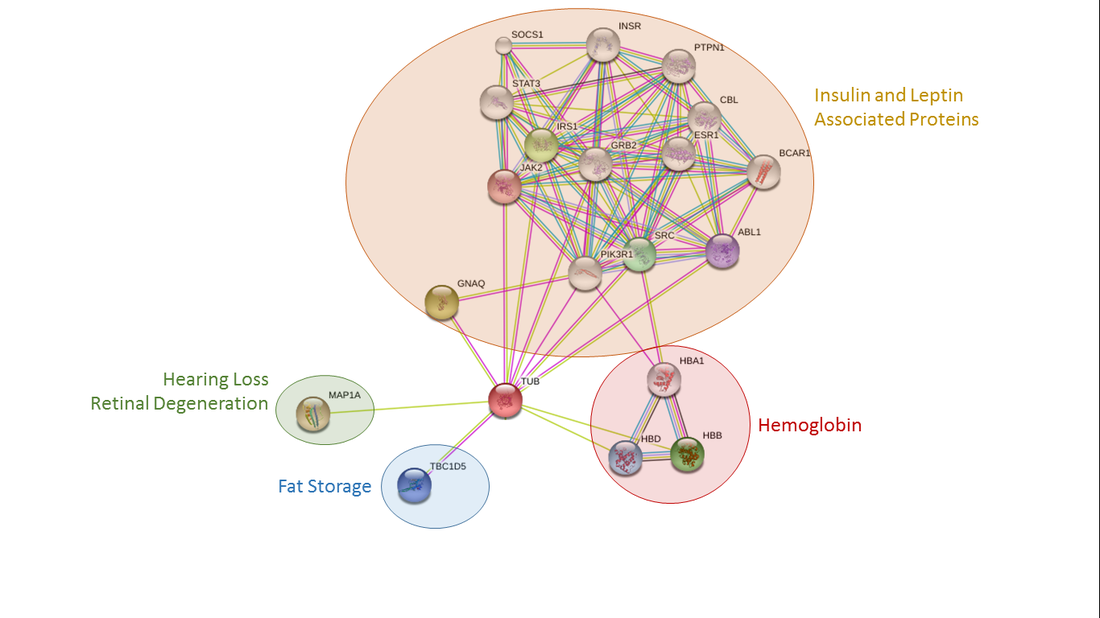
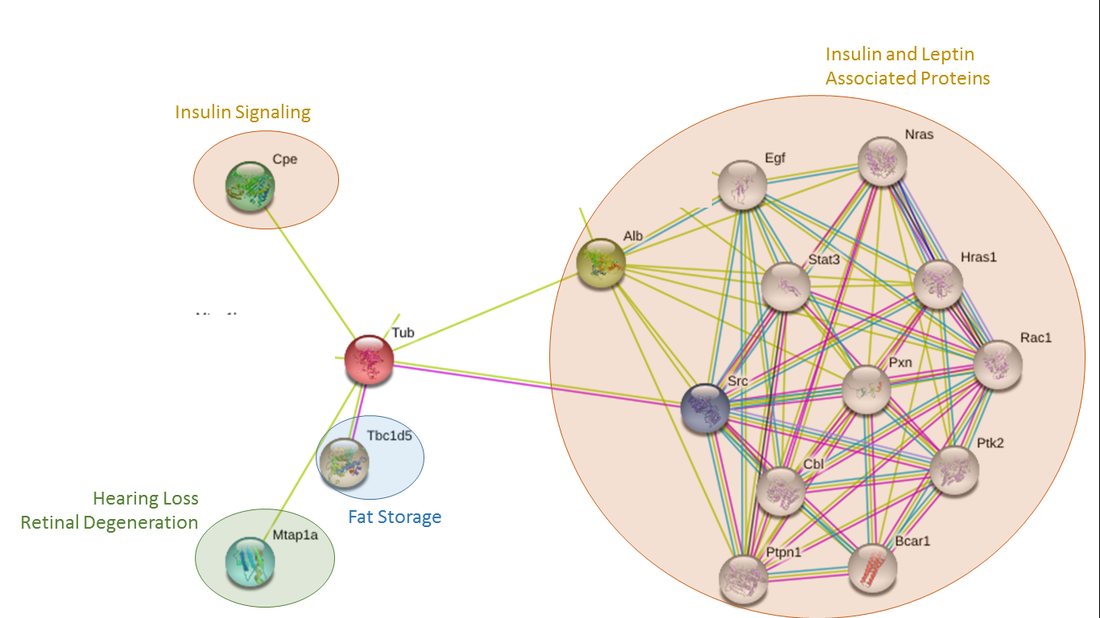
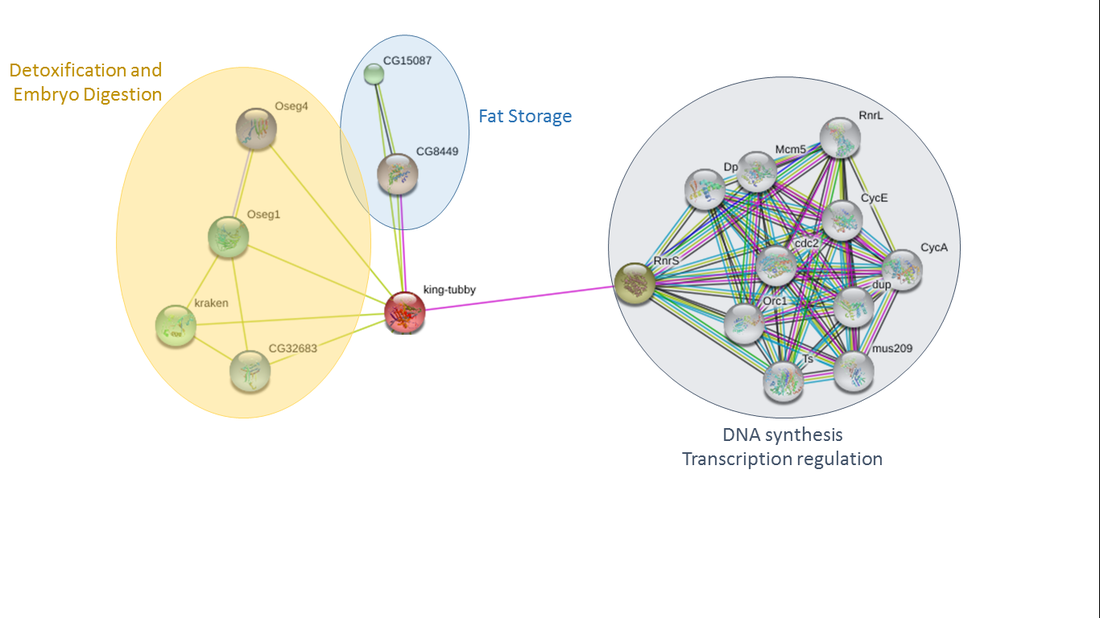
Tubby is predicted to interact with a number of binding partners involved in an array of biological functions. In humans, tubby was predicted to interact with insulin and leptin associated proteins, hemoglobin proteins, proteins involved in fat storage, and a protein involved in hearing loss and retinal degeneration (Figure 1). The same protein groups were identified as interaction partners in mice as well, although no hemoglobin proteins were predicted (Figure 2). Slightly different binding partners were identified for the fruit fly homologue king tubby. King tubby was predicted to interact with fat storage proteins, proteins involved in detoxification and embryo digestion, and DNA synthesis and transcriptional regulation (Figure 3).
analysis of tub protein networks
The protein interaction partners found to interact with tubby in these three organisms support the hypothesis that tubby may function through some interaction with leptin or insulin signaling as previously shown in Prada et al. [2]. In addition, the results in Drosophila are consistent with king tubby acting as a transcription factor. The potential interactions with hemoglobin found in humans, however, is novel and something to be further explored to see if this predicted interaction actually functions in the cell. Future work could focus on verifying these predicted interactions using TAP tags, Y2H screens, and protein pull downs.
REFERENCES
Cover Photo Credit
[1] STRING-Known and Predicted Protein-Protein Interactions, accessed 24 April 2014.
[2] Prada, P.O., Quaresma, P.G.F., Caricilli, A.M., et al. (2013). Tub Has a Key Role in Insulin and Leptin Signaling and Action in Vivo in Hypothalamic Nuclei. Diabetes, 62(1),137, doi: 10.2337/db11-1388.
[1] STRING-Known and Predicted Protein-Protein Interactions, accessed 24 April 2014.
[2] Prada, P.O., Quaresma, P.G.F., Caricilli, A.M., et al. (2013). Tub Has a Key Role in Insulin and Leptin Signaling and Action in Vivo in Hypothalamic Nuclei. Diabetes, 62(1),137, doi: 10.2337/db11-1388.
Site created by Rachael Baird.
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14