This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
The Study of homology
First coined by Richard Owen in 1848, homology is defined as two sequences that share a common ancestor and is fundamental to Charles Darwin's Origin of Species. [1] Since, evolutionary theory predicts that related organisms derived from a common ancestor will share similarities, homology can be used to infer evolutionary relationships. Similarly, evolutionary relationships can be used to understand homology. [2] Gene homologues are also useful in identifying potential model organisms that would allow genetic manipulation and observation not possible in humans as well as indicating the importance and possible function of a gene that has not changed for thousands of years despite other changes throughout the genome. Care must be taken to not confuse analogous organisms with homologous organisms, however. While homologues share a common ancestor and diverge, analogs have separate evolutionary origins but appear superficially similar due to convergent evolution and natural selection. [3]
In a practical sense, it is often impossible to identify every common ancestor since there is no fossil record of these extinct forms. Therefore, a decision on homology between genes has to be made on the basis of the similarity between them and correlated to the probability that this similarity is not simply due to chance. The higher the similarity between two sequences, the lower the probability that they have originated independently of each other and became similar merely by chance. [4]
In a practical sense, it is often impossible to identify every common ancestor since there is no fossil record of these extinct forms. Therefore, a decision on homology between genes has to be made on the basis of the similarity between them and correlated to the probability that this similarity is not simply due to chance. The higher the similarity between two sequences, the lower the probability that they have originated independently of each other and became similar merely by chance. [4]
The homologues of the tub gene
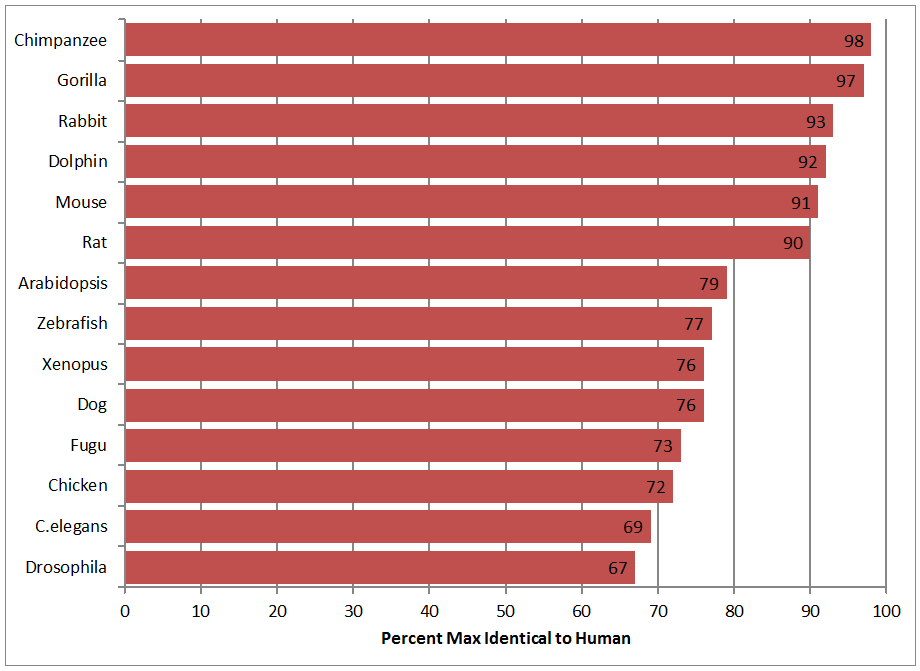
Tub gene homologues were initially identified using NCBI's Basic Local Alignment Search Tool (BLAST). BLAST identifies regions of local similarity between sequences by comparing nucleotide sequences to sequence databases and calculating the statistical significance of the matches. Since Tub is a large gene containing 12 introns, mRNA was used as the input sequence into BLAST rather than genomic DNA to reduce the number of ambiguous alignments occurring in non-coding DNA. After inputting the Tub mRNA sequence, BLAST provided a list of potentially homologous genes in other species. These potential homologues were confirmed through reciprocal BLASTs (using the potential homologue's mRNA as the input sequence to ensure that the human Tub was still identified as a homologue) and sequence alignments to visualize the similarity. Homologene was also used to confirm the homologues initially identified through BLAST. While not all of the identified homologues listed below were found on Homologene, their successful reciprocal BLAST and alignment with the human Tub gene warrants their inclusion as a likely homologue. It is also important to note that the homologues shown in Figure 1 are not an exhaustive list of all potential Tub gene homologues. Instead, the image below displays the great variety of organisms, ranging from plants to humans, that contain Tub gene homologues.
Analysis of Tub Gene homologues
As can be seen in Figure 1, Tub is very well conserved in organisms ranging from plants to flatworms to frogs to chimpanzees, but there were no potential homologues identified in single-cellular organisms, such as yeast. Since all of the organisms shown above were verified to be homologues (and not analogs) through reciprocal BLASTs, it appears that the Tub gene may serve some essential function for all multicellular organisms. This assumption is supported by the fact that the Tub gene has been found to regulate fat storage in mammals as well as C. elegans and Drosophila. [5] These homologues can now be evaluated as potential model organisms for studying tubby.
Homolog reference pages and numbers
|
Human (Homo sapien)-Tub, transcript variant 1
Accession Number: NM_003320.4 GI Number: 222418553 FASTA Gorilla (Gorilla Gorilla)-TUB Accession Number: XM_004050647.1 GI Number: 426367348 FASTA E value: 0.0 Max Identical: 97% Dolphin (Tursiops truncates)-TUB Accession Number: XM_004328674.1 GI Number: 470650709 FASTA E value: 0 Max identical: 92% Rat (Rattus norvegicus)-Tub Accession Number: NM_013077.1 GI Number 6981685 FASTA E value 0 Max identical: 90% Zebrafish (danio rerio)-Tubby homolog (mouse), transcript variant X1 Accession number: XM_005172352.1 GI Number 528484985 FASTA E value: 0 Max identical: 77% Xenopus (Xenopus tropicalis)-tub Accession Number: NM_001007492.1 GI Number: 55926067 FASTA E value: 0 Max identical: 76% Chicken (Gallus gallus)-TUB Accession number: XM_420992.4 GI Number: 513186353 FASTA E value: 0 Max Identical: 72% Drosophila (Drosophila melanogaster)-king tubby Accession Number: NM_206189.1 GI Number 45552772 FASTA E value: 4e-51 Max identical: 67% |
Chimpanzee (Pan troglodytes)-Tub, transcript variant 1
Accession Number: XM_003312904.1 GI Number: 332835807 FASTA E value: 0.0 Max identical: 98% Rabbit (Oryctolagus cuniculus)-Tubby Accession Number: XM_002708791.1 GI Number: 291384589 FASTA E value:0 Max identical: 93% Mouse (Mus musculus) - Tub Accession Number: NM_021885.4 GI Number: 83921622 FASTA E value: 0.0 Max identical: 91% Arabidopsis thaliana-tubby-like F-box protein 3 Accession Number: NM_001202846.1 GI Number: 334184978 FASTA E value: 5e-07 Max Identical: 79% Dog (canis lupus familiaris)-TUB, transcript variant X1 Accession Number: XM_005633625.1 GI Number: 545536700 FASTA E value: 0 Max identical: 76% Fugu (Takifugu rubripes)-tubby protein homolog Accession Number: XM_003970000.1 GI Number: 410913144 FASTA E value: 0.0 Max identical: 73% C. elegans (caenorhabditis elegans)-TUB-1 Accession Number: NM_063309.4 GI Number: 392890781 FASTA E value: 4e-16 Max Identical 69% |
REferences
Cover Photo Credit
[1] Delsuc, F., Brinkmann, H., and Philippe, H. (2005). Phylogenomics and the Reconstruction of the Tree of Life. Nature Reviews Genetics, 6(5), 361. doi: 10.1038/nrg1603.
[2] Understanding Evolution: Homology.
[3] Understanding Evolution: Homologies and Analogies.
[4] Koonin, E.V. and Galperin, M.Y. Sequence-Evolution-Function: Computational Approaches in Comparative Genomics. Boston: Kluwer Academic; 2003. Chapter 2, Evolutionary Concept in Genetics and Genomics. Available from: http://www.ncbi.nlm.nih.gov/books/NBK20255/
[5] Ashrafi K., Chang F.Y., Watts J.L., Fraser A.G., Kamath R.S., Ahringer J., Ruvkun G. (2003). Genome-wide RNAi analysis of Caenorhabditis elegans fat regulatory genes. Nature, 421: 268. doi:10.1038/nature01279.
[1] Delsuc, F., Brinkmann, H., and Philippe, H. (2005). Phylogenomics and the Reconstruction of the Tree of Life. Nature Reviews Genetics, 6(5), 361. doi: 10.1038/nrg1603.
[2] Understanding Evolution: Homology.
[3] Understanding Evolution: Homologies and Analogies.
[4] Koonin, E.V. and Galperin, M.Y. Sequence-Evolution-Function: Computational Approaches in Comparative Genomics. Boston: Kluwer Academic; 2003. Chapter 2, Evolutionary Concept in Genetics and Genomics. Available from: http://www.ncbi.nlm.nih.gov/books/NBK20255/
[5] Ashrafi K., Chang F.Y., Watts J.L., Fraser A.G., Kamath R.S., Ahringer J., Ruvkun G. (2003). Genome-wide RNAi analysis of Caenorhabditis elegans fat regulatory genes. Nature, 421: 268. doi:10.1038/nature01279.
Site created by Rachael Baird.
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14
Genetics 564 Assignment, Spring 2014
University of Wisconsin-Madison
Last Updated: 5-9-14